lesion evolution
Posted: Tue Dec 17, 2019 1:30 am
This article refers to the four possible evolutive types of lesions (active, inactive, chronic active, remyelinating)
Molecular signature of different lesion types in the brain white matter of patients with progressive multiple sclerosis
https://actaneurocomms.biomedcentral.co ... 019-0855-7
► Full pdf
Conclusion
In conclusion, by next-generation RNA sequencing and a comprehensive computational systems medicine approach, we identified the first mechanistic transcriptome signature of lesion evolution and fate in the progressive MS brain WM.
The major de novo network of shared DEGs among the different lesions emphasize inflammation as a common mechanism, and support the view also provided by GWAS that MS is an inflammatory disease, even at the later progressive stage.
We found that each lesion type had a huge complexity of molecular pathways, and although we tried to categorize them to simplify for better understanding, many pathways were unexpected and overlapping indicating a dynamic process of lesion evolution. TGFβR2 was major hub of shared DEGs by all lesions and NAWM, and it was expressed by astrocytes in remyelinating lesions. Our data also suggest that chronic active lesions that are more frequent during the late progressive phase have more complex molecular mechanisms and changes in multiple pathways. This lesion type was profoundly different from all other lesion types, and also from control WM. NAWM was more similar to control WM than to any other lesion types. This indicates that major gene expression changes occur both at early lesion genesis, and in lesions most characteristic as the late progressive phase develops. Besides unique sub-networks mechanistically evolving different lesions stages, some molecules were specifically regulated: CD26/DPP4 by microglia in the NAWM and by mononuclear cells in active lesions. The uniqueness of lesion types also indicates that omics approaches should consider lesion stages, when expression and regulation of different molecules are addressed. Although this study indicates the extreme diverse molecular events on transcriptome level at different lesion stages, our comprehensive unbiased search across subsets of multiple lesions provided a discovery of specific molecular mechanistic signatures validated by different approaches.
A limitation of our study is the absence of controls with other neurological disease, and the lack of separation of rim and core in the chronic active and inactive lesions. Nevertheless, the combination of different bioinformatics methods and validation by immunohistochemistry supported our conclusions, and overcome these limitation for the interpretation of changes in transcriptome signatures during lesion evolution and fate in the WM of progressive MS brain.
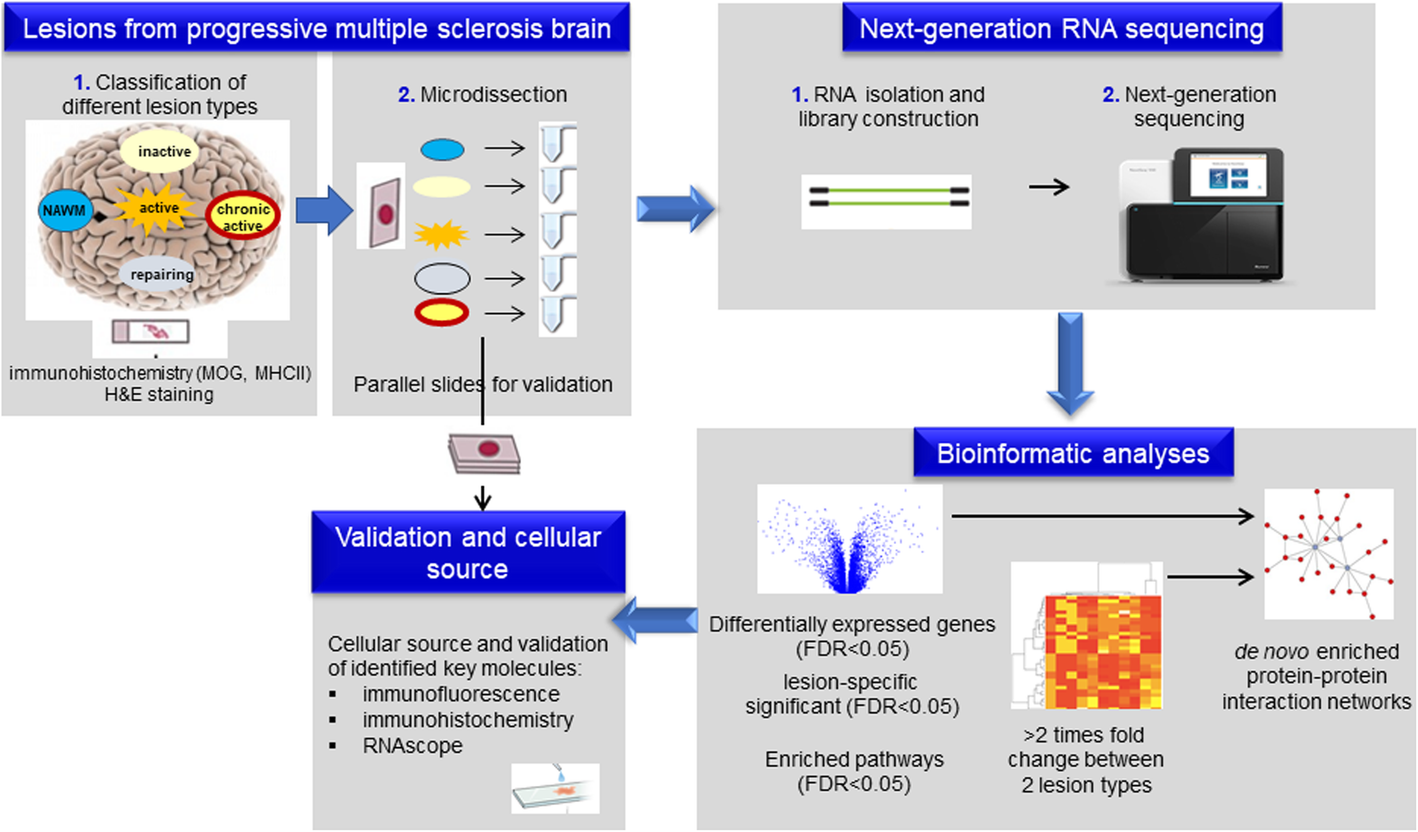
Molecular signature of different lesion types in the brain white matter of patients with progressive multiple sclerosis
https://actaneurocomms.biomedcentral.co ... 019-0855-7
► Full pdf
Conclusion
In conclusion, by next-generation RNA sequencing and a comprehensive computational systems medicine approach, we identified the first mechanistic transcriptome signature of lesion evolution and fate in the progressive MS brain WM.
The major de novo network of shared DEGs among the different lesions emphasize inflammation as a common mechanism, and support the view also provided by GWAS that MS is an inflammatory disease, even at the later progressive stage.
We found that each lesion type had a huge complexity of molecular pathways, and although we tried to categorize them to simplify for better understanding, many pathways were unexpected and overlapping indicating a dynamic process of lesion evolution. TGFβR2 was major hub of shared DEGs by all lesions and NAWM, and it was expressed by astrocytes in remyelinating lesions. Our data also suggest that chronic active lesions that are more frequent during the late progressive phase have more complex molecular mechanisms and changes in multiple pathways. This lesion type was profoundly different from all other lesion types, and also from control WM. NAWM was more similar to control WM than to any other lesion types. This indicates that major gene expression changes occur both at early lesion genesis, and in lesions most characteristic as the late progressive phase develops. Besides unique sub-networks mechanistically evolving different lesions stages, some molecules were specifically regulated: CD26/DPP4 by microglia in the NAWM and by mononuclear cells in active lesions. The uniqueness of lesion types also indicates that omics approaches should consider lesion stages, when expression and regulation of different molecules are addressed. Although this study indicates the extreme diverse molecular events on transcriptome level at different lesion stages, our comprehensive unbiased search across subsets of multiple lesions provided a discovery of specific molecular mechanistic signatures validated by different approaches.
A limitation of our study is the absence of controls with other neurological disease, and the lack of separation of rim and core in the chronic active and inactive lesions. Nevertheless, the combination of different bioinformatics methods and validation by immunohistochemistry supported our conclusions, and overcome these limitation for the interpretation of changes in transcriptome signatures during lesion evolution and fate in the WM of progressive MS brain.
